2. What is Notebook and How to use it
Notebook
Drylab Notebook is a lightweight, stable, notebook-centric environment designed for interactive data exploration, visualization, and standard analyses. It prioritizes simplicity, fast startup, and reproducibility.
This environment is ideal for users who want a clean Jupyter workspace without the overhead of many specialized or experimental packages.
What is the Notebook?
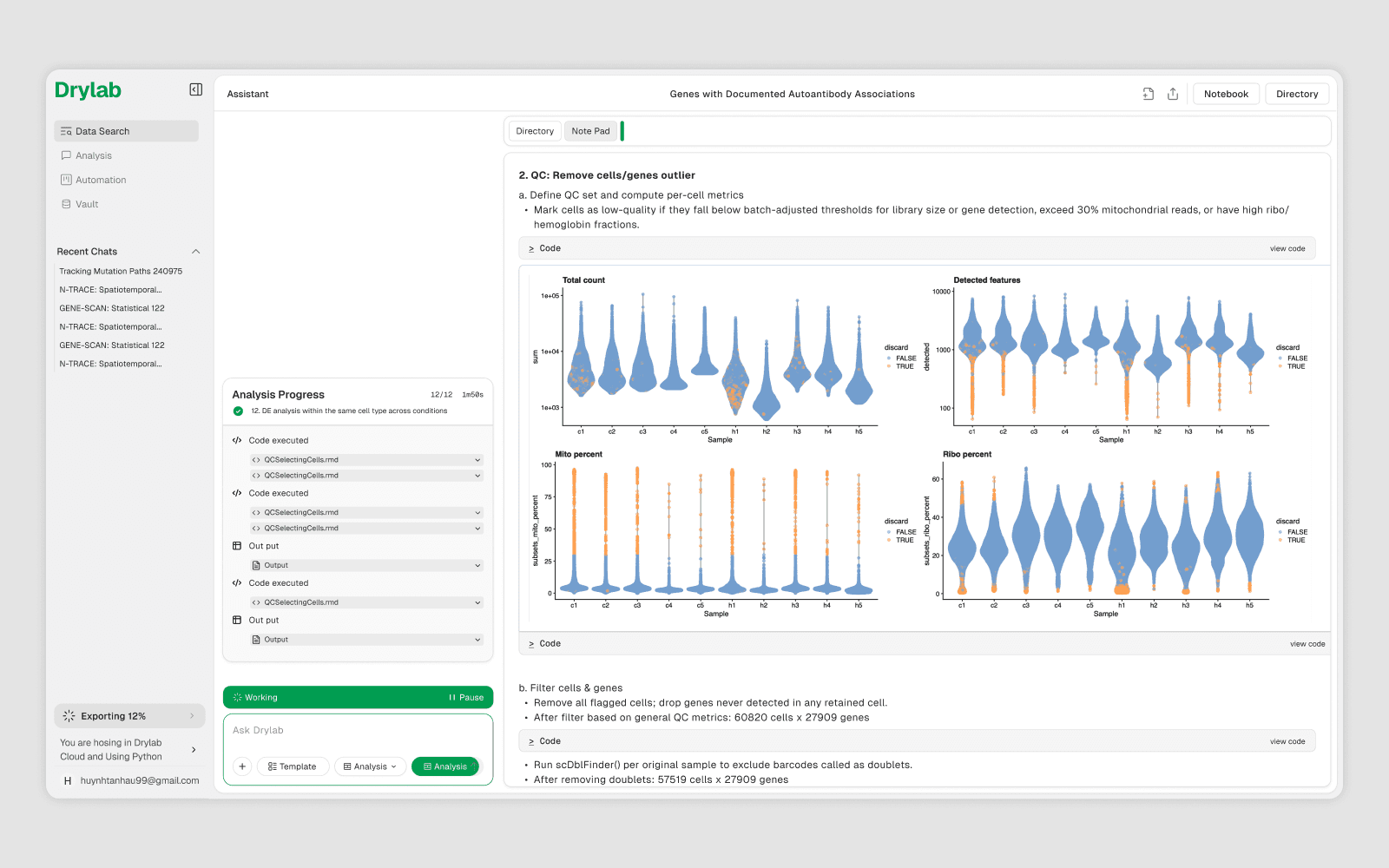
Every analysis session has a persistent notebook — a .ipynb file saved to your workspace. It records every code cell, markdown cell, bash command, and script file generated during the session, making your work reproducible and shareable.
Your notebook location: /Your Project/note.ipynb
Core capabilities
Interactive analysis
Python, R, and Bash notebooks via Jupyter
Step-by-step data exploration
Rapid iteration on analysis and figures
Statistical analysis
Descriptive statistics and hypothesis testing
Regression and classical machine learning
Differential expression for bulk and single-cell data
Light transcriptomic workflows
Basic RNA-seq and scRNA-seq analysis
Clustering, dimensionality reduction, visualization
Simple trajectory and pseudotime analysis
Reporting and visualization
Publication-ready plots
Tables and summary statistics
Shareable notebooks and reports
Included tooling (high level)
Python
Core scientific stack: NumPy, Pandas, SciPy, scikit-learn
Visualization: Matplotlib, Seaborn, Plotly
Transcriptomics: Scanpy, AnnData
Genomics utilities: pysam, Biopython
File formats: HDF5, Parquet, Excel
R
Tidyverse-style data manipulation
Seurat and SeuratObject
Bioconductor core (DESeq2, edgeR, limma)
Single-cell infrastructure (SingleCellExperiment, scater, scran)
Basic trajectory and spatial support (slingshot, SpatialExperiment)
System & runtime
Jupyter Server with Python, R, and Bash kernels
Minimal bioinformatics CLIs (samtools, bedtools, bcftools)
Optimized for stability and low overhead
Typical applications
Exploratory data analysis
Teaching and training
Figure generation for manuscripts
Pilot analyses before scaling up
Users new to computational biology
Cell Types
Type | Purpose | Example |
|---|---|---|
| Run Python (or R) |
|
| Document your work | Headers, notes, captions |
| Shell commands |
|
| Write scripts to disk | R scripts, config files |
Key Notebook Actions
append — Add a new cell at a given index (default for new code)
edit — Modify a cell in-place (simple typo/parameter fixes only)
Ai edit — Modify a cell using AI
edit_and_move — Delete the buggy cell and re-append a fixed version at the end. Use this after debugging, so the notebook runs correctly top-to-bottom.
How to Open Notebook
Click to the Notebook tab on the right corner to open the notebook windows. Toggle between them using the Preview / Edit button at the top of the notebook panel.
Preview Mode
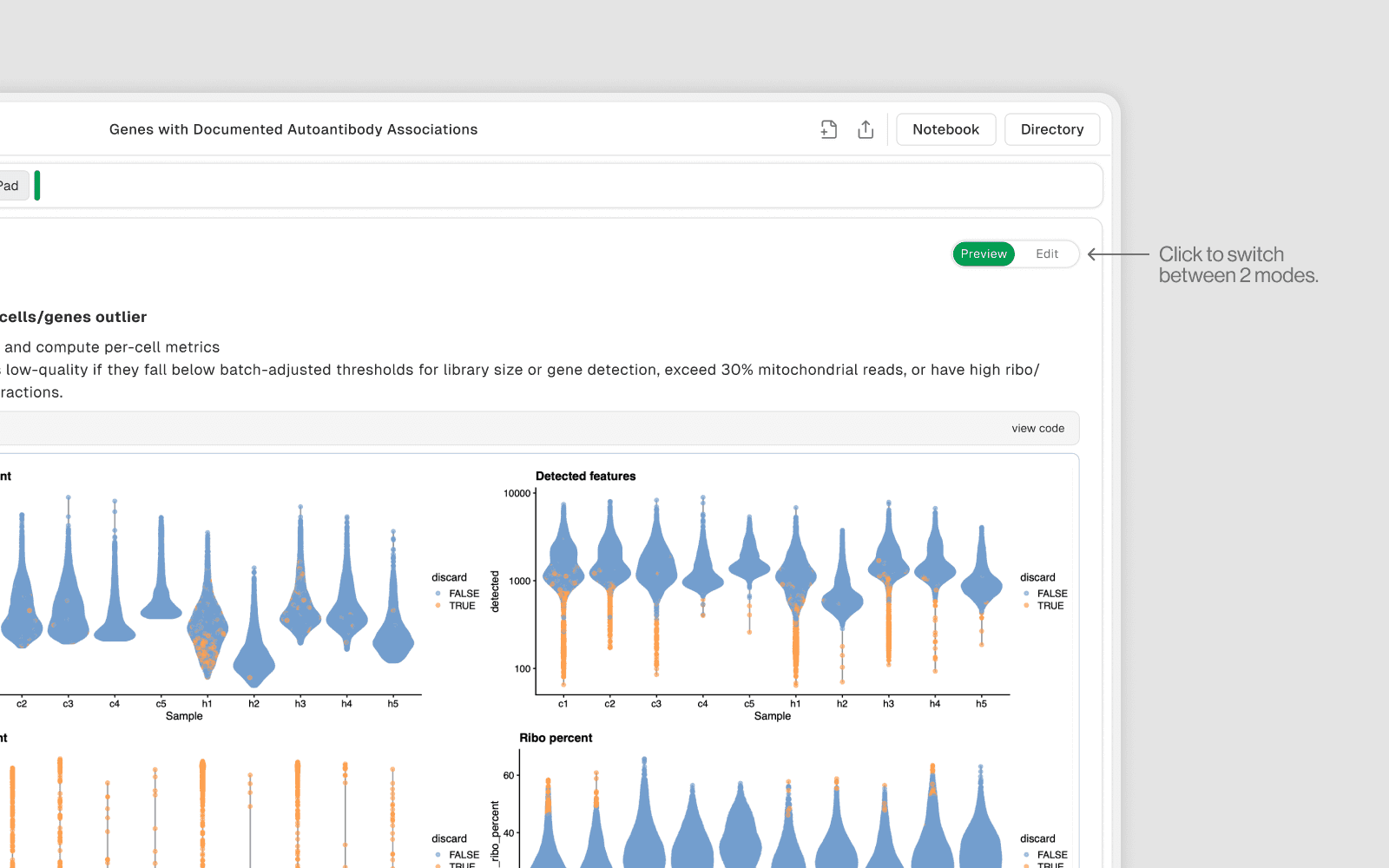
Read-only view of the notebook
Renders markdown cells as formatted text (headers, tables, bold, etc.)
Shows all cell outputs (plots, tables, printed results) from the last run
Use this to review your work, share results, or read documentation
No accidental edits possible
Edit Mode
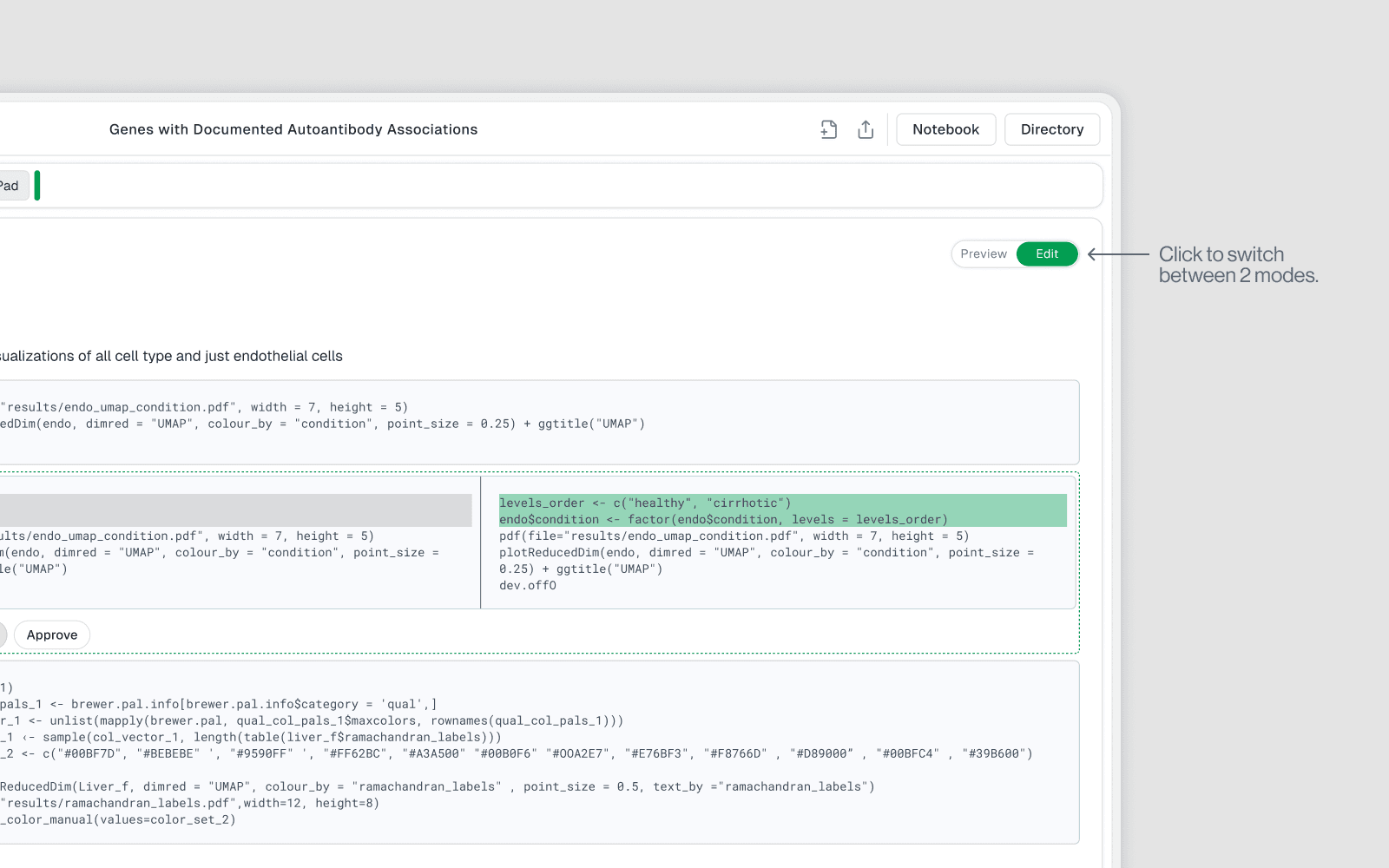
Interactive mode where you can write and modify cells
Markdown cells show raw syntax (e.g.
## Title,*bold**)Code cells are fully editable with syntax highlighting
You can add, remove, reorder, or modify any cell
Use this when actively working on analysis


